NDEC
Overview
NDEC is a hub developed by the Data Bank where neutron data files in the ENDF-6 format are uploaded to. Upon each upload, NDEC automatically performs a sequence of modular processes.
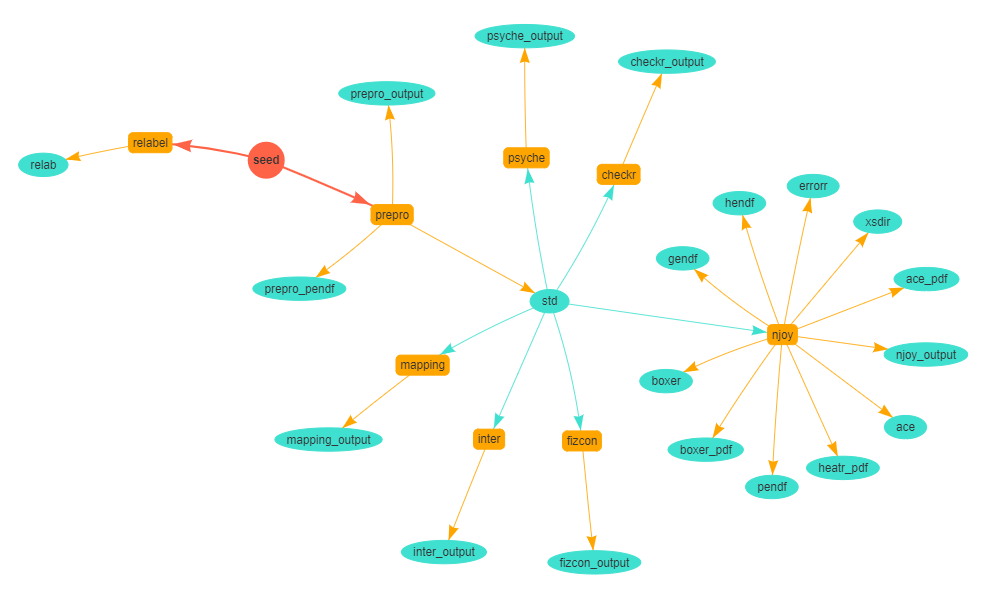
Sequence of processes that are automatically performed by the NDEC system upon uploading an evaluated file or ‘seed’ (in red). Executable scripts are in yellow, the output of these are in blue.
NDEC Modules
NDEC modules that are independent are run in parallel. NDEC modules that need the output of another module as their own input are launched automatically when the needed file becomes available.
| C&U | NJOY-2016 | PREPRO-2017 | NEA |
|---|---|---|---|
| checkr | moder | endf2c | relabel |
| psyche | reconr | linear | libcontent |
| fizcon | broadr | recent | splitting |
| inter | unresr | sigma1 | mapping |
| heatr | legend | benchmarking | |
| gaspr | activate | ||
| thermr | merger | ||
| purr | fixup | ||
| acer | dictin | ||
| groupr | |||
| errorr | |||
| covr |
The final outcome of the NDEC process is an organized structure which the user may access to download processed files, browse the outputs of each module or consult warnings or errors found during the execution of NDEC.
NDEC executes the following codes on each submitted file :
NEA sequences
The NEA Data Bank Nuclear Data Service has developed a series of scripts and data visualization tools to analyse the content of ENDF-6 files and of entire libraries. Below is a list of NEA scripts used in NDEC.
relabel
relabel is a bash script.
relabel is a script that relabels the header section (MF1, MT451) of a file to include the proper library name, version and release date of the file in the ENDF-6 format requirements. It is used for all files in the preparation of a new library release.
libcontent
libcontent is an R script.
libcontent provides an overview of the content of a neutron data library. Three major aspects are covered in its output, which is a webpage for consultation :
Access to all outputs of NDEC run for the library
- Content analysis of the files in the library, in particular :
- Content analysis of all covariance data files in the library, listing all isotopes in the library containing covariance type data, sorted by covariance type (MF)
- Complete overview of the differences that exist in files of a given library (e.g. JEFF-3.3) with respect to another library, which by default is taken as the previous release of the library being analysed (e.g. JEFF-3.2). It provides an exhaustive table of all evaluations that are different in the two libraries.
splitting
splitting is a bash script.
splitting an ENDF-6 file is breaking it up into its different MF/MT sections. The files is parsed to look for any of the ENDF-6 format dividers (SEND, FEND, MEND, TEND).Before this is done, the evaluated file is standardized with the code endf2c.
mapping
mapping is an R script.
Mapping an ENDF-6 file is building a graphical representation of the different parts (MF/MT sections) that constitute the evaluated file, and how these sections match others from other library releases. It is also a visual way of finding out which type of information is contained in a given file.
Below is an example of the JEFF-3.3 U-235 map, where the different colors relate to matches found for in other libraries.
In order to find which sections match which, a one-to-one comparison of each MF/MT standardized section is performed through a database of MF/MT sections. The map reports the oldest ‘match’ that was found. When a section is reported as matching the library the file is coming from, it means this section was last updated or added in the release considered.
The figure below gives the list of all releases considered in this database.
